Improving the Effectiveness of the Annual Flu Vaccine

Table of Contents
Author(s)
Kirstin R.W. Matthews
Fellow in Science and Technology PolicyShare this Publication
- Download PDF
- Print This Publication
- Cite This Publication Copy Citation
Deem, Michael W., Melia E. Bonomo, and Kirstin R.W. Matthews. 2018. Improving the Effectiveness of the Annual Flu Vaccine. Policy brief no. 10.29.18. Rice University’s Baker Institute for Public Policy, Houston, Texas.
Seasonal influenza (flu) causes 9.3 million to 49.0 million illnesses and 12,000 to 79,000 deaths in the United States annually.1 To combat its impact, the Centers for Disease Control and Prevention (CDC) recommends all healthy children and adults obtain a flu vaccination every year. In 2017-18, 57.9% of healthy children (6 months to 17 years old) obtained the vaccine, and approximately 80% of pediatric deaths from influenza during that season were children who were unvaccinated.2 Vaccinations have been proven to reduce flu-related hospitalizations and severe illness outcomes.3,4
Figure 1 — Flu Vaccination Coverage and Vaccine Effectiveness Against Flu Type A(H3N2) for U.S. Adults
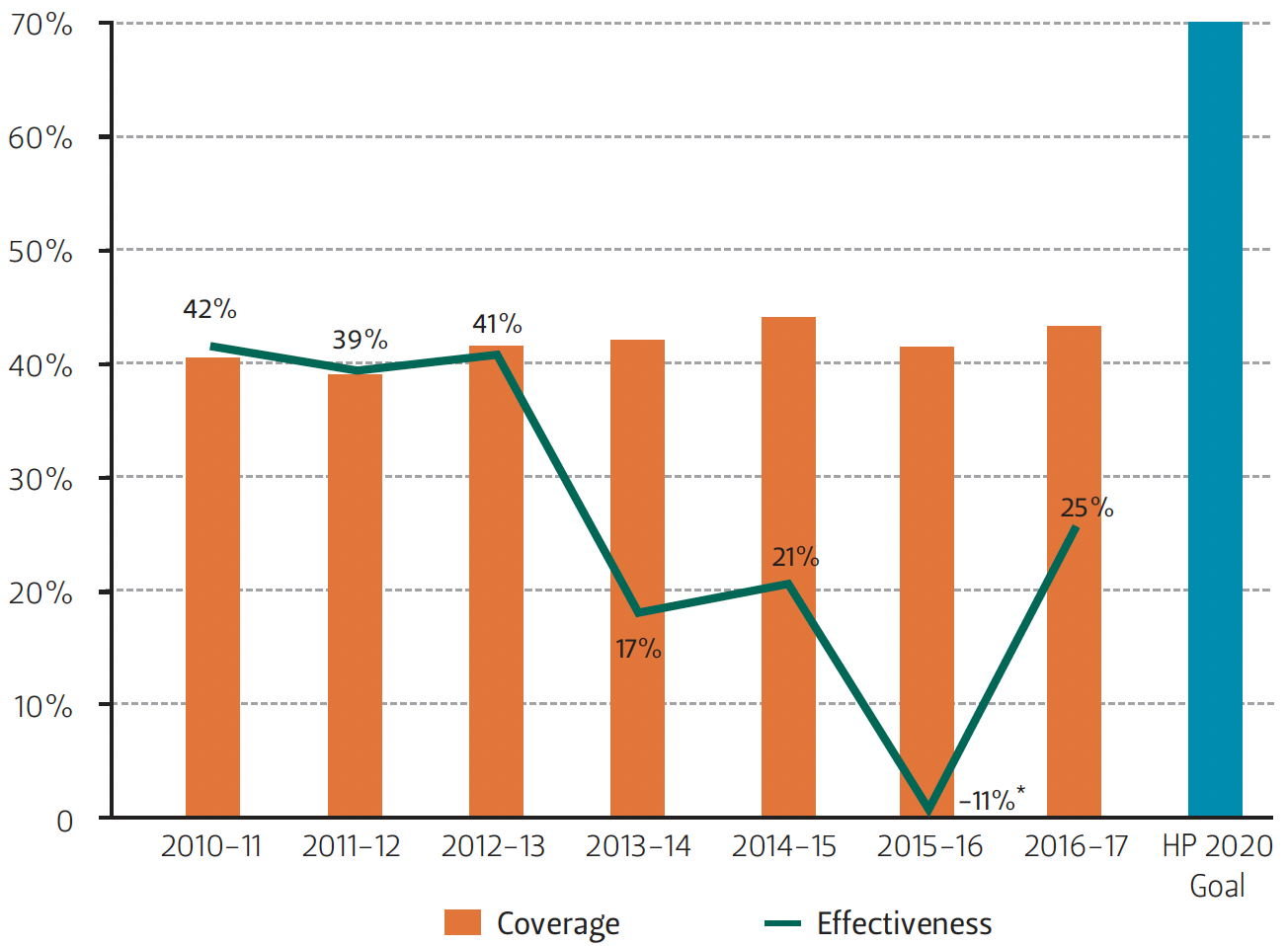
*In 2015-16, the flu vaccination effectiveness against type A(H3N2) was -11%, which means that people who received the vaccine were more susceptible to being infected with the A(H3N2) flu virus than people who were unvaccinated.
Source [Coverage] Centers for Disease Control and Prevention (CDC), https://www.cdc.gov/flu/fluvaxview/coverage-1617estimates.htm; [Effectiveness] Bonomo, M.E., and M.W. Deem. 2018. Clinical Infectious Diseases 67(7): 1129–1131.
One challenge to flu vaccinations is that a new flu vaccine is usually created each year. The flu virus is constantly evolving and changing.5 The genetic material of the viruses is encoded in RNA, an unstable molecule that is characterized by frequent mutations, or random changes to the genome. Because of these genetic changes, the vaccine must be updated almost every flu season.5,6 Predicting the most dominant flu virus strains that will affect people each flu season is a difficult process, as demonstrated by the variability in the effectiveness of flu vaccines (Figure 1).
The majority of flu vaccines are created from three types of deactivated flu viruses: A(H3N2), A(H1N1), and B. The A(H3N2) virus is usually the most dominant and causes the majority of patient illness and death each flu season because it tends to mutate more frequently than the other types.7 Over the past decade, the A(H3N2) vaccine reduced the risk of having to go to the doctor with the flu by 25% (Figure 1).8
To develop a vaccine in time for the beginning of flu season in the fall, scientists must start in early January. The current method that the CDC uses involves scientists vaccinating ferrets with several vaccine candidates. They then extract the antibodies from the ferrets to estimate which vaccine was the most effective against the dominant viruses from the previous flu season. This method has been used for almost 50 years. However, it has been proven to be inconsistent in predicting how well the vaccines would perform in humans, especially with the recent, rapidly mutating A(H3N2) viruses.7-9 Additionally, experiments with ferrets are time-consuming and costly.
In contrast, mathematical models, including a model developed at Rice University over 15 years ago, allow scientists to calculate how well the flu vaccine matches the infecting viruses.10,11 The Rice model, called pEpitope, estimates vaccine effectiveness, and it has been shown to work well for flu A(H3N2), A(H1N1), and B vaccines.8,12,13 For the 2018-19 flu season, Rice scientists predict that the vaccine will be between 20–40% effective against the majority of A(H3N2) viruses. Unfortunately, the model also predicts no protection against a small, newly evolving group of A(H3N2) flu strains. Modeling, if used during the early stages of vaccine development, also helps identify likely vaccine strains and improves the effectiveness of the vaccine.
Recommendations
Public health researchers are often slow to change. Despite the fact that Rice’s pEpitope model has been around for more than 15 years, it is unclear why the CDC has yet to take advantage of it in developing their seasonal flu vaccine.10, 11
Adding such a model to the already existing ferret experiments will enhance the current vaccination decision-making process. This mathematical modeling technique can rapidly narrow down the viruses that would be good candidates for the vaccine during a particular flu season. It can serve as a check to make sure that the vaccine virus does not mutate during the manufacturing process. The pEpitope model is also low-cost, as it does not require any specialized equipment. Finally, it is extremely quick, taking just a couple of seconds to analyze the potential effectiveness of a vaccine against thousands of infecting viruses in a particular geographic region.
The CDC should strengthen their current protocols for choosing vaccine candidates by utilizing all available prediction modeling. This will improve overall flu vaccine effectiveness rates and potentially coverage rates as well. Scientists hope that with improved effectiveness, they will also be able to improve vaccine coverage rates, which still trail behind the CDC’s Healthy People 2020 goal of 70%.14 Given the difficulty in producing effective vaccines and the general climate of public mistrust of immunization, this work has the potential to improve vaccine strain selection and education by providing a tool that is accessible to researchers and citizen scientists alike.
Endnotes
1. Centers for Disease Control and Prevention (CDC). 2018. “Disease Burden of Influenza.” https://www.cdc.gov/flu/about/disease/burden.htm.
2. Centers for Disease Control and Prevention (CDC). 2018. “Cumulative monthly influenza vaccination coverage estimates for children 6 months through 17 years by state, HHS region, and the United States, National Immunization Survey- Flu (NIS-Flu), 2017-18 influenza season.” https://www.cdc.gov/flu/fluvaxview/reportshtml/reporti1718/reporti/index.html.
3. Talbot, H.K., et al. 2013. “Effectiveness of influenza vaccine for preventing laboratory-confirmed influenza hospitalizations in adults, 2011–2012 influenza season.” Clinical Infectious Diseases 56(12): 1774–1777.
4. Arriola, C., et al. 2017. “Influenza vaccination modifies disease severity among community-dwelling adults hospitalized with influenza.” Clinical Infectious Diseases 65(8): 1289–1297.
5. Petrova, V.N., and C.A. Russell. 2018. “The evolution of seasonal influenza viruses.” Nature Reviews Microbiology 16(1): 47.
6. Palese, P. 2004. “Influenza: old and new threats.” Nature Medicine 10(12s): S82.
7. Hampson, A., et al. 2017. “Improving the selection and development of influenza vaccine viruses–Report of a WHO informal consultation on improving influenza vaccine virus selection, Hong Kong SAR, China, 18–20 November 2015.” Vaccine 35(8): 1104–1109.
8. Bonomo, M.E., and M.W. Deem. 2018. “Predicting Influenza H3N2 Vaccine Efficacy from Evolution of the Dominant Epitope.” Clinical Infectious Diseases 67(7): 1129–1131.
9. Barr, I.G., et al. 2010. “Epidemiological, antigenic and genetic characteristics of seasonal influenza A (H1N1), A (H3N2) and B influenza viruses: basis for the WHO recommendation on the composition of influenza vaccines for use in the 2009–2010 Northern Hemisphere season.” Vaccine 28(5): 1156–1167.
10. Deem, M.W., and H.Y. Lee. 2003. “Sequence Space Localization in the Immune System Response to Vaccination and Disease.” Physical Review Letters 91: 068101.
11. Morris, D.H., et al. 2018. “Predictive Modeling of Influenza Shows the Promise of Applied Evolutionary Biology." Trends in Microbiology 26(2): 102-118.
12. Pan, K., K.C. Subieta, and M.W. Deem. 2010. “A novel sequence-based antigenic distance measure for H1N1, with application to vaccine effectiveness and the selection of vaccine strains.” Protein Engineering, Design & Selection 24(3): 291–299.
13. Pan, Y., and M.W. Deem. 2016. “Prediction of influenza B vaccine effectiveness from sequence data.” Vaccine 34(38): 4610–4617.
14. Centers for Disease Control and Prevention (CDC). 2018. “Results for General Population Influenza Vaccination Coverage.” https://www.cdc.gov/flu/fluvaxview/interactive-general-population.htm.
This material may be quoted or reproduced without prior permission, provided appropriate credit is given to the author and Rice University’s Baker Institute for Public Policy. The views expressed herein are those of the individual author(s), and do not necessarily represent the views of Rice University’s Baker Institute for Public Policy.


